MAKA-Mmap——Prediction of Protein Real-Valued distance
Presentations
MAKA-Mmap provides an online tool for real-valued distance prediction between protein residues. Users can upload pre-extracted feature files in .pkl format for inference and visualization. The following notes may be helpful:
· MAKA-Mmap only accepts .pkl files containing protein residue features in a predefined format. Please ensure that the file meets the required structure before uploading. Only one file can be uploaded at a time.
· After submission, the system will return a .npy file containing the predicted real-valued distance map, along with a visualized heatmap for easier interpretation.
· Prediction time may vary depending on the sequence length and complexity of the input.
· You may click on pre-loaded examples to quickly test the prediction functionality without uploading your own file.
· If you encounter any issues or have questions, feel free to contact us via the email provided in the top-right corner.
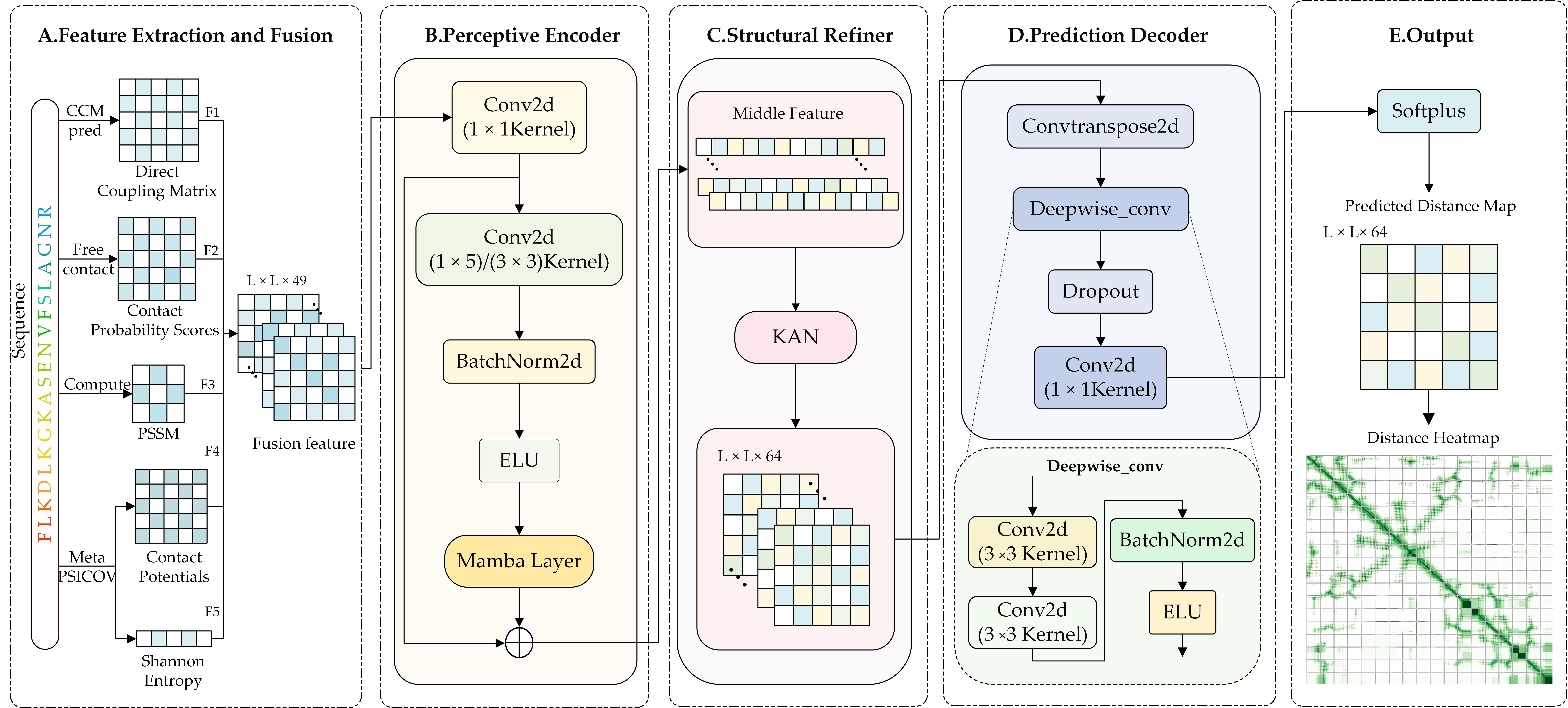
· The figure below illustrates the conceptual design and output of the MAKA-Mmap framework.